
Contingency and selection in mitochondrial genome dynamics
High frequencies of mutant mitochondrial DNA (mtDNA) in human cells lead to cellular defects that are associated with aging and disease. Yet much remains to be understood about the dynamics of the generation of mutant mtDNAs and their relative replicative fitness that informs their fate within cells and tissues. To address this, we utilize long-read single-molecule sequencing to track mutational trajectories of mtDNA in the model organism Saccharomyces cerevisiae. This model has numerous advantages over mammalian systems due to its much larger mtDNA and ease of artificially competing mutant and wild-type mtDNA copies in cells. We show a previously unseen pattern that constrains subsequent excision events in mtDNA fragmentation in yeast.
Current Member of this project: Chris Nunn

Behavior of Competitive Ecosystems
Ecological communities are shaped by the inter- and intra-species interactions. We examine how these interactions together with other factors, e.g., immigration, control the community’s composition and long-term dynamics in the presence of noise and fluctuations. We address this problem using a minimal null model of interacting multi-species ecological communities that incorporates competition, immigration, and demographic noise. Interestingly, we find that a complete phase diagram exhibits rich behavior with multiple regimes that go beyond the classical ‘niche’ and ‘neutral’ regimes. We study the transitions between regimes, their characteristic dynamics and species-species correlation.
Current Member of this project: Nava Leibovich, Sid Goyal in collaboration with Jeremy Rothschild and Anton Zilman.
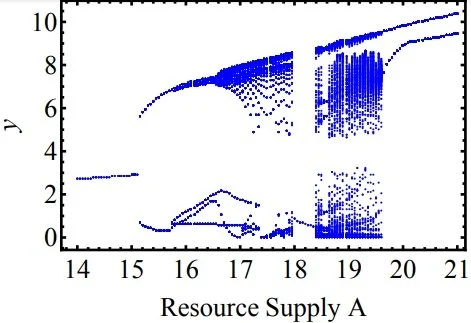
Ecosystem Diversity
In ecosystems where competition is fierce, how does diversity develop against Darwinian’s survival of the fittest? One of the plausible approaches is that the fitness landscape may be flattened by self-organization of a combination of strains in the system, under constraints or trade-offs imposed on metabolism. Using coarse-grained models on metabolism, our work bridges the scales on a cellular level with that on a population level and shows how self-organization leads to formation of niche, which effectively changes the environment to lower inter-strain competition, thereby equalizing differences in fitness. In large systems, this is related to connectivity and network motifs, and may provide new ways to quantify diversity in experiments
Current Member of this project: Sam Lui

Diversity-generating host-disease coevolution with adaptive immunity
Bacteria are under constant threat from viruses, and some bacteria possess adaptive immune systems that provide protection through a genomic 'memory' of past viral infections. In conjunction with viral evolution, this creates a diverse population of bacteria where each cell has a unique viral genomic imprint. The fate of the bacterial population depends on the rate at which memories are updated to track evolving viruses. How does the dynamic fitness landscape generated by adaptive immunity impact population diversity and the rate of evolution? We find with a simple stochastic population dynamics model that viruses experience a changing fitness that both drives and reigns in diversity: new virus mutants that escape immune targeting have high fitness and experience selective dynamics associated with low diversity, while established virus clones experience immune targeting and lose their fitness advantage, going extinct following neutral dynamics associated with high diversity.
Current Member of this project: Madeleine Bonsma-Fisher

C. Elegans Escape Response
Under the co-supervision of Dr. Sid Goyal and Dr. Will Ryu, I am pursuing a research project studying changes in the escape response of C. elegans as they age. This study involves lasering the worms in the head and precisely tracking their escape attempt as shown in the provided figure. The aim is to demonstrate that a behavioral multi-system test can describe the health of C. elegans well, and that such a description will more completely elucidate the changes to their health through aging than more time-consuming lifespan studies.
Current Member of this project: Colins Brandy
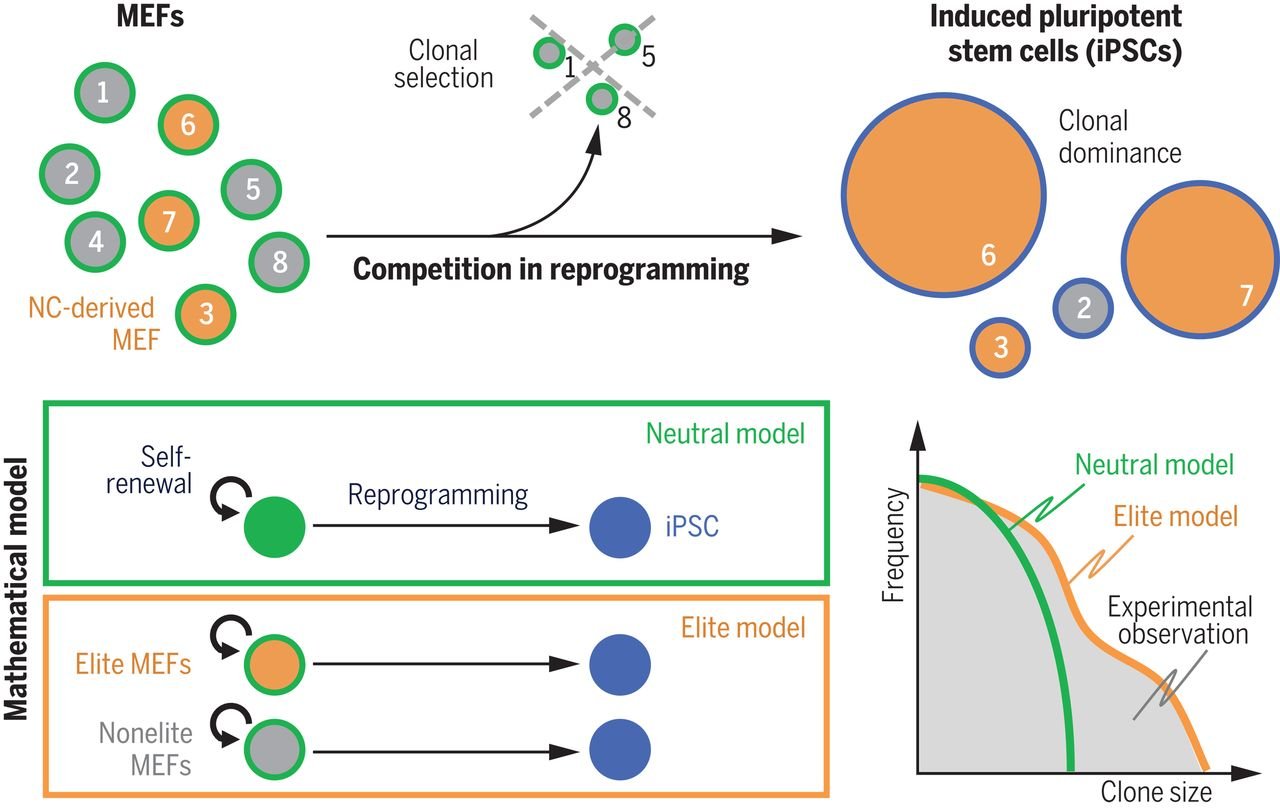
Competition in Reprogramming
A Nobel Prize–winning discovery showed that specialized cells can be genetically reprogrammed into stem cells, thus gaining the ability to become any cell type in the body. But what happens during reprogramming is not completely understood. Shakiba et al. used experimental and mathematical approaches to show that skin cells compete during reprogramming, eliminating one another as the population progresses toward the stem cell state (see the Perspective by Wolff and Purvis). The “winners” are a special class of skin cells originating from the neural crest. Cells of this type normally emerge during embryonic development and migrate into various tissues, including the skin, muscle, and nervous system.
Current Member of this project: Sophie McGibbons-Gardner